# Run this cell to set up packages for lecture.
from lec18_imports import *
Agenda¶
- Recap: The Central Limit Theorem (CLT).
- Choosing sample sizes.
- Models.
Recap: The Central Limit Theorem¶
show_clt_slides()
The Central Limit Theorem¶
- The Central Limit Theorem (CLT) says that the probability distribution of the sum or mean of a large random sample drawn with replacement will be roughly normal, regardless of the distribution of the population from which the sample is drawn.
- The distribution of the sample mean is centered at the population mean, and its standard deviation is
$$\text{SD of Distribution of Possible Sample Means} = \frac{\text{Population SD}}{\sqrt{\text{sample size}}}$$
- When we collect a single sample, we won't know the population mean or SD, so we'll use the sample mean and SD as approximations.
- Key idea: The CLT allows us to estimate the distribution of the sample mean, given just a single sample, without having to bootstrap!
Concept Check ✅ – Answer at cc.dsc10.com¶
Which one of these histograms corresponds to the distribution of the sample mean for samples of size 100 drawn from a population with mean 50 and SD 20?
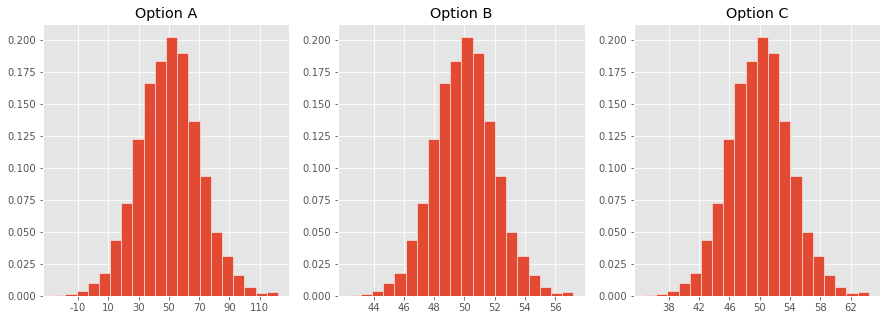
Confidence intervals for the population mean¶
A 95% confidence interval for the population mean is given by
$$ \left[\text{sample mean} - 2\cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}}, \text{sample mean} + 2\cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}} \right] $$
This CI doesn't require bootstrapping, and it only requires three numbers – the sample mean, the sample SD, and the sample size!
Bootstrapping vs. the CLT¶
Bootstrapping still has its uses!
| Bootstrapping | CLT | |
|---|---|---|
| Pro | Works for many sample statistics (mean, median, standard deviation). |
Only requires 3 numbers – the sample mean, sample SD, and sample size. |
| Con | Very computationally expensive (requires drawing many, many samples from the original sample). |
Only works for the sample mean (and sum). |
Choosing sample sizes¶
Polling¶
- You want to estimate the proportion of UCSD students that use Duolingo, which is the most popular language learning app.

- To do so, you will ask a random sample of UCSD students whether or not they use Duolingo.
- You want to create a confidence interval that has:
- A 95% confidence level.
- A width of at most 0.06.
- The interval (0.21, 0.25) would be fine, but the interval (0.21, 0.28) would not.
Question: How big of a sample do you need? 🤔
Aside: Proportions are means!¶
- The sample we collect will consist of only two unique values:
- 1, if the student uses Duolingo.
- 0, if they don't.
- We're interested in the proportion of values in our sample that are 1.
- This proportion is the same as the mean of our sample!
- For instance, suppose our sample is $0, 1, 1, 0, 1$. Then $\frac{3}{5}$ of the values are $1$. The sample mean is
$$\frac{0 + 1 + 1 + 0 + 1}{5} = \frac{3}{5}$$
Key takeaway: The CLT applies in this case as well! The distribution of the proportion of 1s in our sample is roughly normal.
Our strategy¶
We will:
- Collect a random sample.
- Compute the sample mean (i.e., the proportion of people who say "yes").
- Compute the sample standard deviation.
- Construct a 95% confidence interval for the population mean:
$$ \left[ \text{sample mean} - 2\cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}}, \ \text{sample mean} + 2\cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}} \right] $$
Note that the width of our CI is the right endpoint minus the left endpoint:
$$ \text{width} = 4 \cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}} $$
Our strategy¶
- We want a CI whose width is at most 0.06.
$$\text{width} = 4 \cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}}$$
- The width of our confidence interval depends on two things: the sample SD and the sample size.
- If we know the sample SD, we can find the appropriate sample size by re-arranging the following inequality:
$$4 \cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}} \leq 0.06$$
$$\sqrt{\text{sample size}} \geq 4 \cdot \frac{\text{sample SD}}{0.06} \\ \implies \boxed{\text{sample size} \geq \left( 4 \cdot \frac{\text{sample SD}}{0.06} \right)^2}$$
- Problem: Before polling, we don't know the sample SD, because we don't have a sample! We don't know the population SD either.
- Solution: Find an upper bound – i.e. the largest possible value – for the sample SD and use that.
Upper bound for the standard deviation of a sample¶
- Without any information about the values in a sample, we can't make any guarantees about the standard deviation of the sample.
- However, in this case, we know that the only values in our sample will be 0 ("no") and 1 ("yes").
- In Homework 5, we introduce a formula for the standard deviation of a collection of 0s and 1s:
$$\text{SD of Collection of 0s and 1s} = \sqrt{(\text{Prop. of 0s}) \times (\text{Prop. of 1s})}$$
# Plot the SD of a collection of 0s and 1s with p proportion of Os.
p = np.arange(0, 1.01, 0.01)
sd = np.sqrt(p * (1 - p))
plt.plot(p, sd)
plt.xlabel('p')
plt.ylabel(r'$\sqrt{p(1-p)}$');
- Fact: The largest possible value of the SD of a collection of 0s and 1s is 0.5.
- This happens when half the values are 0 and half are 1.
Choosing a sample size¶
- In the sample we will collect, the maximum possible SD is 0.5.
- Earlier, we saw that to construct a confidence interval with the desired confidence level and width, our sample size needs to satisfy:
$$\text{sample size} \geq \left( 4 \cdot \frac{\text{sample SD}}{0.06} \right)^2$$
- Notice that as the sample SD increases, the required sample size increases.
- By using the maximum possible SD above, we ensure that we collect a large enough sample, no matter what the population and sample look like.
Choosing a sample size¶
$$\text{sample size} \geq \left( 4 \cdot \frac{\text{sample SD}}{0.06} \right)^2$$
By substituting 0.5 for the sample size, we get
$$\text{sample size} \geq \left( 4 \cdot \frac{\text{0.5}}{0.06} \right)^2$$
While any sample size that satisfies the above inequality will give us a confidence interval that satisfies the necessary properties, it's time-consuming to gather larger samples than necessary. So, we'll pick the smallest sample size that satisfies the above inequality.
(4 * 0.5 / 0.06) ** 2
1111.1111111111113
Conclusion: We must sample 1112 people to construct a 95% CI for the population mean that is at most 0.06 wide.
Activity¶
Suppose we instead want a 95% CI for the population mean that is at most 0.03 wide. What is the smallest sample size we could collect?
✅ Click here to see the answer after you've attempted the question yourself.
$\text{sample size} \geq \left( 4 \cdot \frac{\text{0.5}}{0.03} \right)^2 = 4444.44$ Therefore, the smallest sample size we could collect is 4445.
Statistical models¶
Statistical inference¶
- So far in the second half of this class, we've focused on the problem of parameter estimation.
- Given a single sample, we can construct a confidence interval using bootstrapping (for most statistics) or the CLT (for the sample mean).
- This confidence interval gives us a range of estimates for a parameter.
- Next, we'll turn our attention to answering yes-no questions about the relationships between samples and populations.
- Example: Does it look like this jury panel was drawn randomly from this population of eligible jurors?
- Example: Does it look like this sequence of coin tosses was generated by a fair coin?
- Both of these problems fall under the umbrella of statistical inference – using a sample to draw conclusions about the population.
Models¶
A model is a set of assumptions about how data was generated.

Goal¶
- Our goal is to assess the quality of a model.
- Suppose we have access to a dataset. What we'll try to do is determine whether a model "explains" the patterns in the dataset.
Example: Jury selection¶
Swain vs. Alabama, 1965¶
- Robert Swain was a Black man convicted of crime in Talladega County, Alabama.
- He appealed the jury's decision all the way to the Supreme Court, on the grounds that Talladega County systematically excluded Black people from juries.
- At the time, only men 21 years or older were allowed to serve on juries. 26% of this eligible population was Black.
- But of the 100 men on Robert Swain's jury panel, only 8 were Black.
Supreme Court ruling¶
- About disparities between the percentages in the eligible population and the jury panel, the Supreme Court wrote:
"... the overall percentage disparity has been small...”
- The Supreme Court denied Robert Swain’s appeal and he was sentenced to life in prison.
- We now have the tools to show quantitatively that the Supreme Court's claim was misguided.
- This "overall percentage disparity" turns out to be not so small, and is an example of racial bias.
- Jury panels were often made up of people in the jury commissioner's professional and social circles.
- Of the 8 Black men on the jury panel, none were selected to be part of the actual jury.
Setup¶
- Model: Jury panels consist of 100 men, randomly chosen from a population that is 26% Black.
- Observation: On the actual jury panel, only 8 out of 100 men were Black.
- Question: Does the model explain the observation?
Our approach: Simulation¶
- We'll start by assuming that the model is true.
- We'll generate many jury panels using this assumption.
- We'll count the number of Black men in each simulated jury panel to see how likely it is for a random panel to contain 8 or fewer Black men.
- If we see 8 or fewer Black men often, then the model seems reasonable.
- If we rarely see 8 or fewer Black men, then the model may not be reasonable.
- Run an experiment once to generate one value of our chosen statistic.
- In this case, sample 100 people randomly from a population that is 26% Black, and count the number of Black men (statistic).
- Run the experiment many times, generating many values of the statistic, and store these statistics in an array.
- Visualize the resulting empirical distribution of the statistic.
Step 1 – Running the experiment once¶
- How do we randomly sample a jury panel?
np.random.choicewon't help us, because we don't know how large the eligible population is.
- The function
np.random.multinomialhelps us sample at random from a categorical distribution.
np.random.multinomial(sample_size, pop_distribution)
np.random.multinomialsamples at random from the population, with replacement, and returns a random array containing counts in each category.pop_distributionneeds to be an array containing the probabilities of each category.
Aside: Example usage of np.random.multinomial
On Halloween 👻, you trick-or-treated at 35 houses, each of which had an identical candy box, containing:
- 30% Starbursts.
- 30% Sour Patch Kids.
- 40% Twix.
At each house, you selected one candy blindly from the candy box.
To simulate the act of going to 35 houses, we can use np.random.multinomial:
np.random.multinomial(35, [0.3, 0.3, 0.4])
array([ 8, 14, 13])
Step 1 – Running the experiment once¶
In our case, a randomly selected member of our population is Black with probability 0.26 and not Black with probability 1 - 0.26 = 0.74.
demographics = [0.26, 0.74]
Each time we run the following cell, we'll get a new random sample of 100 people from this population.
- The first element of the resulting array is the number of Black men in the sample.
- The second element is the number of non-Black men in the sample.
np.random.multinomial(100, demographics)
array([35, 65])
Step 1 – Running the experiment once¶
We also need to calculate the statistic, which in this case is the number of Black men in the random sample of 100.
np.random.multinomial(100, demographics)[0]
26
Step 2 – Repeat the experiment many times¶
- Let's run 10,000 simulations.
- We'll keep track of the number of Black men in each simulated jury panel in the array
counts.
counts = np.array([])
for i in np.arange(10000):
new_count = np.random.multinomial(100, demographics)[0]
counts = np.append(counts, new_count)
counts
array([21., 32., 32., ..., 31., 31., 31.])
Step 3 – Visualize the resulting distribution¶
Was a jury panel with 8 Black men suspiciously unusual?
(bpd.DataFrame().assign(count_black_men=counts)
.plot(kind='hist', bins = np.arange(9.5, 45, 1),
density=True, ec='w', figsize=(10, 5),
title='Empiricial Distribution of the Number of Black Men in Simulated Jury Panels of Size 100'));
observed_count = 8
plt.axvline(observed_count, color='pink', linewidth=4, label='Observed Number of Black Men in Actual Jury Panel')
plt.legend();
# In 10,000 random experiments, the panel with the fewest Black men had how many?
counts.min()
11.0
Conclusion¶
- Our simulation shows that there's essentially no chance that a random sample of 100 men drawn from a population in which 26% of men are Black will contain 8 or fewer Black men.
- As a result, it seems that the model we proposed – that the jury panel was drawn at random from the eligible population – is flawed.
- There were likely factors other than chance that explain why there were only 8 Black men on the jury panel.
Summary, next time¶
Summary¶
- A 95% confidence interval for the population mean is given by $$ \left[\text{sample mean} - 2\cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}}, \text{sample mean} + 2\cdot \frac{\text{sample SD}}{\sqrt{\text{sample size}}} \right] $$
- If we want to construct a confidence interval of a particular width and confidence level for a population proportion:
- Choose a confidence level (e.g. 95%) and maximum width (e.g. 0.06).
- Solve for the minimum sample size that satisfies both conditions.
- Collect a sample of that size.
- Use the formula above to construct an interval.
- A model is an assumption about how data was generated. We're interested in determining the validity a model, given some data we've collected.
- When assessing a model, we consider two viewpoints of the world: one where the model is true, and another where the model is false for some reason.
Next time¶
- Next time, we'll see more examples of testing models and deciding between two viewpoints.
- We'll formalize this notion, which is called hypothesis testing.